[Researcher Comment] Whole-genome sequencing helps halt MRSA bacterial outbreak
Image: CC
Dr. William Hanage, PhD and Associate Professor at the Harvard School of Public Health, talks to 2 Minute Medicine:
 “Without question this sort of approach is the way forward. The investigators have been nothing short of visionary in how they have brought modern tech to the bedside. You can generate these sorts of data even now for a few tens of [dollars] per isolate. That is only going to fall until the sequence of an infecting bug will be another item in a medical record, like BP or BMI. This is almost without question what molecular epidemiology is going to look like in the future.
“Without question this sort of approach is the way forward. The investigators have been nothing short of visionary in how they have brought modern tech to the bedside. You can generate these sorts of data even now for a few tens of [dollars] per isolate. That is only going to fall until the sequence of an infecting bug will be another item in a medical record, like BP or BMI. This is almost without question what molecular epidemiology is going to look like in the future.
The ‘almost’ in that sentence refers to a concern, that I share, that our ability to obtain sequence is lagging behind our ability to interpret it.”
Key study points:
- Whole-genome sequencing was able to identify cases in an outbreak more accurately than standard methods and enabled the identification of a complex transmission network spanning both the hospital and community.
- Whole-genome sequencing provided reliable data in a clinically relevant time frame, enabling useful interventions.
Primer: The ability to track and trace a disease outbreak is critical to controlling and perhaps even ending it. John Snow famously demonstrated this when he identified the Broad Street pump as the source of the 1854 London cholera epidemic. Armed with this knowledge, he implemented a public health intervention that halted further spread of the disease – he had the pump handle removed.
The outbreak investigation toolkit has expanded somewhat since John Snow’s day. Nowadays, laboratory techniques allow discrimination between different strains of a pathogen, enabling finer resolution of transmission networks. These techniques include antibiotic-susceptibility profiles (antibiograms) and molecular methods like pulsed-field gel electrophoresis and multi-locus sequence typing.
Ultimately, the various strains of a pathogen differ based on their underlying genetic makeup. Consequently, strain typing methods can be organized on a continuum of genetic resolution. Antibiotic susceptibility, for example, is a coarse indicator of genetic composition; it provides indirect evidence for what antibiotic resistance genes a strain does or does not have. Pulsed-field gel electrophoresis deals more directly with the actual genetic material; it distinguishes strains using the pattern of bands that their DNA makes after being cut up and run on a gel. Multi-locus sequence typing looks even more closely at strains’ genetic compositions. In this method, several designated locations in the genome are sequenced and strains are distinguished by the combination of sequences they possess at each of these locations.
On this continuum, the ultimate in genetic resolution is whole-genome sequencing. As the price, speed and reliability of sequencing technology continue to improve, novel and fruitful applications of whole-genome sequencing will continue to arise.
Background reading:
- Coverage of the study in Science
- Coverage of the study in Nature
- Blog post by Prof. Ed Feil critiquing the study, with response by study author Julian Parkhill in the comments
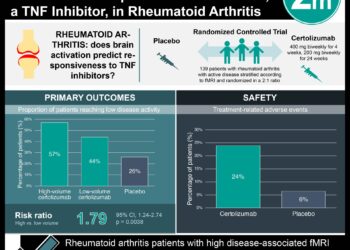
This [descriptive] study: An alleged methicillin resistant Staph aureus (MRSA) outbreak was detected in the special care baby unit (SCBU) of Cambridge University Hospitals. The infection control team used antibiograms to determine that 12 of 17 MRSA isolates during a 6 month period were related. However, they were unable to conclude that there had been an ongoing outbreak because of gaps of several weeks when there were no cases in the unit.
Whole-genome sequencing (WGS) revealed that 14 of the 17 isolates were in fact related; 2 isolates had been wrongly classified by antibiograms. By performing WGS on other MRSA isolates from elsewhere in the hospital, the authors were able to construct a complex transmission network involving mothers of babies in the unit as well as spouses.
The authors also screened 154 SCBU staff and found one staff member who was an asymptomatic carrier of ST2371. The staff member was successfully decolonized.
In sum: This study is novel in its application of whole-genome sequencing to an ongoing outbreak. It is also a powerful demonstration of the capability of existing WGS technology to provide useful data in a clinically relevant timeframe and at a reasonable cost.
Compared to conventional methods, WGS enabled more accurate case identification and facilitated the identification of a complex transmission network. The authors were also able to identify a staff member who may have acted as a reservoir for the pathogen during periods with no cases.
The sequence data generated by WGS also present novel opportunities for analysis. For example, the authors suggest that the sequence diversity of strains found on the colonized staff member can be used to back-calculate the length of time the staffer had been colonized. Some question whether this can truly be done in the context of the present study (see #3 in Background Reading), but they also acknowledge that WGS data from outbreaks open new avenues for research and analysis.
Click to read the study in The Lancet Infectious Diseases
By [TJ] and [MP]
© 2012 2minutemedicine.com. All rights reserved. No works may be reproduced without written consent from 2minutemedicine.com. Disclaimer: We present factual information directly from peer reviewed medical journals. No post should be construed as medical advice and is not intended as such by the authors or by 2minutemedicine.com. PLEASE SEE A HEALTHCARE PROVIDER IN YOUR AREA IF YOU SEEK MEDICAL ADVICE OF ANY SORT.