Genetic variations may predict outcomes for colorectal cancer patients
Image: PD
1. Single nucleotide polymorphisms (SNPs) in LKB1, AMPK, and TSC1 genes were identified as predictive of decreased patient survival.
2. A greater number of high-risk alleles per patient was correlated with decreased survival when compared to patients with zero high-risk alleles.
Evidence Rating Level: 2 (Good)
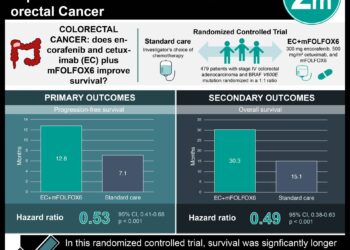
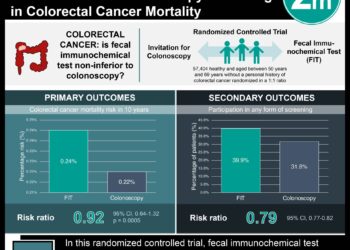
Study Rundown: Nearly 5% of Americans will be diagnosed with colorectal cancer in their lives, resulting in approximately 16.4 deaths per 100,000 people (NCI SEER). The primary treatment for colorectal cancer is surgical resection of the tumor. Patient characteristics including age and tumor stage are often used as predictors of patient outcome and are applied to determine post-surgery therapy regimes including adjuvant chemotherapy. Postulating that an increased number of predictive markers for patient outcomes should lead to more effect post-surgery treatment plans, this study searched for genetic markers to predict colorectal cancer patient outcomes. mTOR activity, which is regulated through the LKB1-AMPK-mTOR pathway, is frequently disregulated in cancer cells and is believed to, in part, drive the cancer phenotype. This study evaluated patient tissue samples for genetic variations within this pathway in search of predictive markers of disease progression. Three particular SNPs, genetic variants in the LKB1, AMPK and TSC1 genes, were found to significantly correlate with decreased disease free survival (DFS) and overall survival (OS). Additionally, the number of these three SNPs (1, 2 or 3) per patient was negatively correlated with survival. These findings suggest that in addition to anatomical and pathological patient characteristics, genetic tests for these particular biomarkers may provide greater insight into colorectal patient outcomes. This study was limited by its multiple comparisons methodology and did not consider the variation in adjuvant therapies in their multivariate analysis.
Click to read the study in the Annals of Surgical Oncology
Relevant Reading: Genetic variation in a metabolic signaling pathway and colon and rectal cancer risk: mTOR, PTEN, STK11, RPKAA1, PRKAG2, TSC1, TSC2, PI3K and Akt1
In-Depth [prospective cohort]: This study examined patient samples for 18 SNPs within 6 genes previously associated with colorectal cancer. Patient tissue samples were collected from 772 Korean patients during tumor resection surgery between 2001 and 2008. Patient outcomes were subsequently documented. SNP genotyping completed via MassARRAY assay and genotype were grouped according to a dominant and recessive model. The Kaplan-Meier method was used to obtain survival estimates and Cox’s proportional hazards model was used for multivariate analysis, accounting for age, preoperative carcinoembryonic antigen and tumor stage. Three genes were identified as significantly correlated with patient DFS and/or OS (p < 0.05): LKB1 (STK11), AMPK (PRKAA1), and TSC1. Specific genetic variations, or SNPs, within these three genes were found to drive these correlations. In particular, three SNPs (STK11 rs741765 GG genotype, PRKAA1 rs461404 TT genotype, and TSC1 rs13295634 TG/GG genotype) were correlated with patient survival no mater primary site or node involvement. Patient survival was also dependent on the number (1, 2 or 3) of these particular SNP alleles when compared with subjects that had zero of these high-risk SNPs (OS HR = 1.629, 2.730 and 3.111; p = 0.135, 0.002, 0.003, respectively).
More from this author: Neuroendocrine tumor marker pancreastatin may be predictive of survival, Initial kidney transplant factors may provide insight into retransplant success
©2012-2014 2minutemedicine.com. All rights reserved. No works may be reproduced without expressed written consent from 2minutemedicine.com. Disclaimer: We present factual information directly from peer reviewed medical journals. No post should be construed as medical advice and is not intended as such by the authors, editors, staff or by 2minutemedicine.com. PLEASE SEE A HEALTHCARE PROVIDER IN YOUR AREA IF YOU SEEK MEDICAL ADVICE OF ANY SORT.