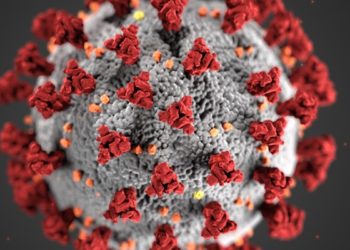
Novel coronavirus identified from patients with pneumonia in Wuhan, China
1. According to an investigation by a rapid response team, the Wuhan outbreak was most likely caused by a novel betacoronavirus.
2. This study reinforces the utility of next-generation sequencing and bioinformatics, especially in the identification of poorly-characterized pathogens.
Evidence Rating Level: 2 (Good)
Study Rundown: Coronaviruses, RNA-containing particles enveloped with protruding glycoproteins, are widely distributed in mammal and avian populations. In December of 2019, several cases of pneumonia with unknown cause were identified in Wuhan, the capital of Hebei province in China. An investigation launched by the Chinese CDC detected a novel strain of coronavirus (2019-nCov) that was traced back to a wet animal seafood market. Lower respiratory tract samples were collected from patients who presented with pneumonia of unknown cause as well as control patients with pneumonia of known cause. Following extraction of nucleic acids from these samples, viral RNA was isolated and sequenced, yielding an 87% match with a previously published bat SARS-like coronavirus genome. Virion particles were found to be generally spherical with ~10 nm spikes via transmission electron microscopy, and cytopathic effects were observed in airway epithelial cells 4 days after inoculation. While this study did not fulfil Koch’s postulates, evidence of seroconversion and pathogenicity have helped etiologically implicate 2019-nCov in the disease outbreak. Further research is needed to determine the coronavirus’s modes of transmission and develop effective strategies to control its spread.
Click here to read the study in NEJM
Relevant Reading: Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR
In-Depth [case series]: In this study designed to identify the cause of viral pneumonia in clusters of patients in Wuhan, China, various molecular techniques, high-throughput sequencing, and transmission electron microscopy were utilized to detect, isolate, and visualize a novel sarbecovirus. After collecting lower respiratory tract samples, including bronchoalveolar-lavage fluid, nucleic acids were extracted and tested for foreign material using polymerase chain reaction (PCR) and a real-time reverse transcription PCR (RT-PCR) assay. Supernatant from bronchoalveolar-lavage fluid samples was inoculated on special-pathogen-free human epithelial cell cultures and monitored daily with light microscopy and RT-PCR. Cells that showed cytopathic effects were negative-stained and subjected to transmission electron microscopy; virion particles were found to be generally spherical with some pleomorphism and had an average diameter of 60 to 140 nm with prominent spikes characteristic of the coronaviridae family. Reads generated from Illumina and nanopore sequencing were assembled into contig maps, and genome gaps were filled using rapid amplification of cDNA ends. From these, three complete genome sequences were crafted, revealing a 86.9% nucleotide sequence identity to a known bat SARS-like Cov genome. Additional evidence supporting the discovery of a novel coronavirus include identification of a specific antigen through immunohistochemical analysis and confirmation of pathogenicity through monkey experiments.
Image: PD
©2020 2 Minute Medicine, Inc. All rights reserved. No works may be reproduced without expressed written consent from 2 Minute Medicine, Inc. Inquire about licensing here. No article should be construed as medical advice and is not intended as such by the authors or by 2 Minute Medicine, Inc.

![Macitentan better than placebo for preventing progression of pulmonary arterial hypertension [SERAPHIN Trial]](https://www.2minutemedicine.com/wp-content/uploads/2013/08/ca68dc0329bef0b42ccae7b6656a0f_gallery-350x250.jpg)

![ABCD2 Score: Predicting Early Stroke Risk After Transient Ischemic Attack (TIA) [Classics Series]](https://www.2minutemedicine.com/wp-content/uploads/2013/05/web-cover-classics-with-logo-medicine-BW-small-jpg-350x250.jpg)